|
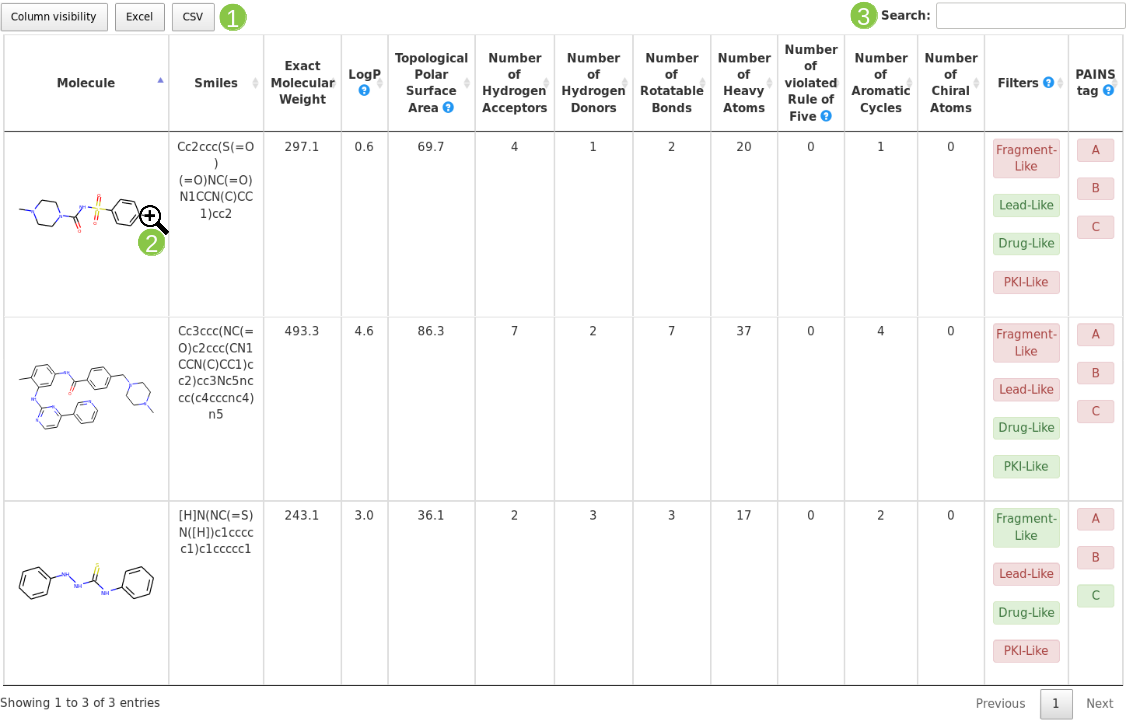
Filtering Critieria |
Comments/Citation |
| Fragment-like |
LogP <=3.5 and MW <=250 and NRB <=5 |
Carr, Congreve, Murray and Rees, Drug Discovery Today, 2005 |
| Lead-like |
MW <= 350 and MW>= 250 and LogP <= 3.5 and NRB <=7 |
Teague, Davis, Leeson and Oprea, Angewandte Chemie International Edition, 1999 |
| Drug-like |
MW <= 500 and MW>= 150 and LogP <= 5 and NRB <=7 and TPSA < 150 and HBD <=5 and HBA <=10 |
Lipinski, Journal of Pharmacological and Toxicological Methods, 2000 |
| PKI-like |
1 <= NAR <= 5 and NCA <=2. No more than 2 values out of the following range : 314 <= MW <= 613, 55 <= TPSA <= 138, 3 <= HBA <= 10, HBD <= 4, 1 <= NRB <=11 and 0.7 <= ClogP <= 6.3. |
Carles, Bourg, Meyer and Bonnet, Molecules, 2018 ; Bournez,
Carles, Peyrat, Aci-Sèche, Bourg, Meyer and Bonnet, Molecules, 2020 |
| GSK 4/400 |
LogP < 4 and MW < 400 |
Gleeson, Journal of Medicinal Chemistry, 2008 |
| Pfizer 3/75 |
LogP > 3 and TPSA < 75 |
Hughes, Blagg, Price, Bailey, DeCrescenzo, Devraj, Ellsworth, Fobian, Gibbs, Gilles, Greene, Huang, Krieger-Burke, Loesel, Wager, Whiteley, Zhang, Bioorganic & Medicinal Chemistry Letters, 2008 |
| PAINS |
Filters A, B and C were extracted from the original study of Jonathan B. Baell and Georgina A. Holloway
|
Baell and Holloway, Journal of Medicinal Chemistry, 2010
Greg Landrum, RDKit blogspot, 2015
|